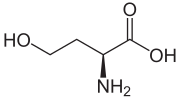
Homoserine
| |

| |
| Names | |
|---|---|
| IUPAC name
(S)-2-Amino-4-hydroxybutanoic acid
| |
| Identifiers | |
3D model (JSmol)
|
|
| ChEBI | |
| ChEMBL | |
| ChemSpider | |
| ECHA InfoCard | 100.010.538 |
| EC Number |
|
PubChem CID
|
|
| UNII | |
CompTox Dashboard (EPA)
|
|
| |
| |
| Properties | |
| C4H9NO3 | |
| Molar mass | 119.12 g/mol |
| Melting point | 203 °C (decomposes) |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |
Homoserine (also called isothreonine) is an
Homoserine is an intermediate in the biosynthesis of three essential amino acids: methionine, threonine (an isomer of homoserine), and isoleucine.[1] Its complete biosynthetic pathway includes glycolysis, the tricarboxylic acid (TCA) or citric acid cycle or the Krebs cycle, and the aspartate metabolic pathway. It forms by two reductions of aspartic acid via the intermediacy of aspartate semialdehyde.I[2] Specifically, the enzyme homoserine dehydrogenase, in association with NADPH, catalyzes a reversible reaction that interconverts L-aspartate-4-semialdehyde to L-homoserine. Then, two other enzymes, homoserine kinase and homoserine o-succinyl transferase use homoserine as a substrate and produce phosphohomoserine and o-succinyl homoserine respectively.[3]
Applications
Commercially, homoserine can serve as precursor to the synthesis of isobutanol and 1,4-butanediol.[4] Purified homoserine is used in enzyme structural studies.[5] Also, homoserine has played important roles in studies to elucidate peptide synthesis and synthesis of proteoglycan glycopeptides.[6] Bacterial cell lines can make copious amounts of this amino acid.[3][4]
Biosynthesis
Homoserine is produced from aspartate via aspartate-4-semialdehyde, which is produced from

L-Homoserine is substrate for homoserine kinase, yielding phosphohomoserine (homoserine-phosphate), which is converted to by threonine synthase to yield L-threonine.
Homoserine is converted to o-succinyl homoserine by homoserine o-succinyl transferase, a precursor to L-methionine.[8]
Homoserine allosterically inhibits aspartate kinase and glutamate dehydrogenase.[3] Glutamate dehydrogenase reversibly converts glutamate to
References
- ^ Tanaka M, Kishi T, Kinoshita S (September 1961). "Studies on the Synthesis of l -Amino Acids: Part III. A Synthesis of l -Homoserine from l -Aspartic Acid". Agricultural and Biological Chemistry. 25 (9): 678–679. doi:10.1080/00021369.1961.10857862. ISSN 0002-1369.
- ^ Berg, J. M.; Stryer, L. et al. (2002), Biochemistry. W.H. Freeman. ISBN 0-7167-4684-0
- ^ a b c Liu P, Zhang B, Yao ZH, Liu ZQ, Zheng YG (October 2020). Zhou NY (ed.). "Multiplex Design of the Metabolic Network for Production of l-Homoserine in Escherichia coli". Applied and Environmental Microbiology. 86 (20). Bibcode:2020ApEnM..86E1477L. doi:10.1128/AEM.01477-20. PMC 7531971. PMID 32801175.
- ^ a b Huang JF, Zhang B, Shen ZY, Liu ZQ, Zheng YG (July 2018). "Metabolic engineering of E. coli for the production of O-succinyl-l-homoserine with high yield". 3 Biotech. 8 (7): 310. doi:10.1007/s13205-018-1332-x. PMC 6037649. PMID 30002999.
- ^ Akai S, Ikushiro H, Sawai T, Yano T, Kamiya N, Miyahara I (February 2019). "The crystal structure of homoserine dehydrogenase complexed with l-homoserine and NADPH in a closed form". Journal of Biochemistry. 165 (2): 185–195. doi:10.1093/jb/mvy094. PMID 30423116.
- ^ Yang W, Ramadan S, Yang B, Yoshida K, Huang X (December 2016). "Homoserine as an Aspartic Acid Precursor for Synthesis of Proteoglycan Glycopeptide Containing Aspartic Acid and a Sulfated Glycan Chain". The Journal of Organic Chemistry. 81 (23): 12052–12059. doi:10.1021/acs.joc.6b02441. PMC 5215661. PMID 27809505.
- ^ Viola, Ronald E. (2001). "The Central Enzymes of the Aspartate Family of Amino Acid Biosynthesis". Accounts of Chemical Research. 34 (5): 339–349. doi:10.1021/ar000057q. PMID 11352712.
- ^ a b Petit C, Kim Y, Lee SK, Brown J, Larsen E, Ronning DR, et al. (January 2018). "Reduction of Feedback Inhibition in Homoserine Kinase (ThrB) of Corynebacterium glutamicum Enhances l-Threonine Biosynthesis". ACS Omega. 3 (1): 1178–1186. doi:10.1021/acsomega.7b01597. PMC 6045374. PMID 30023797.
