Abstract
Hutchinson–Gilford progeria syndrome (HGPS) is a rare lethal genetic disorder characterized by symptoms reminiscent of accelerated aging. The major underlying genetic cause is a substitution mutation in the gene coding for lamin A, causing the production of a toxic isoform called progerin. Here we show that reduction of lamin A/progerin by a single-dose systemic administration of adeno-associated virus-delivered CRISPR–Cas9 components suppresses HGPS in a mouse model.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
9,800 Yen / 30 days
cancel any time
Subscription info for Japanese customers
We have a dedicated website for our Japanese customers. Please go to natureasia.com to subscribe to this journal.
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Data availability
The accession number for the RNA-Seq data reported in this paper is NCBI GEO: GSE122865.
References
Worman, H. J., Fong, L. G., Muchir, A. & Young, S. G. J. Clin. Invest. 119, 1825–1836 (2009).
Broers, J. L., Ramaekers, F. C., Bonne, G., Yaou, R. B. & Hutchison, C. J. Physiol. Rev. 86, 967–1008 (2006).
Kubben, N. & Misteli, T. Nat. Rev. Mol. Cell Biol. 18, 595–609 (2017).
Gordon, L. B., Rothman, F. G., Lopez-Otin, C. & Misteli, T. Cell 156, 400–407 (2014).
Ahmed, M. S., Ikram, S., Bibi, N. & Mir, A. Mol. Neurobiol. 55, 4417–4427 (2018).
Gonzalo, S., Kreienkamp, R. & Askjaer, P. Ageing Res. Rev. 33, 18–29 (2017).
Gordon, L. B. et al. Circulation 130, 27–34 (2014).
Gordon, L. B. et al. GeneReviews® [Internet] (NCBI Bookshelf, 2003).
Young, S. G., Yang, S. H., Davies, B. S., Jung, H. J. & Fong, L. G. Sci. Transl. Med. 5, 171ps3 (2013).
Vidak, S. & Foisner, R. Histochem. Cell. Biol. 145, 401–417 (2016).
De Sandre-Giovannoli, A. et al. Science 300, 2055 (2003).
Eriksson, M. et al. Nature 423, 293–298 (2003).
Buchwalter, A. & Hetzer, M. W. Nat. Commun. 8, 328 (2017).
Osorio, F. G. et al. Sci. Transl. Med. 3, 106ra107 (2011).
Fong, L. G. et al. J. Clin. Invest. 116, 743–752 (2006).
Lopez-Mejia, I. C. et al. EMBO Rep. 15, 529–539 (2014).
Olaya Santiago-Fernández, F. G. O. et al. Nat. Med. (in the press).
Ocampo, A. et al. Cell 167, 1719–1733 e12 (2016).
Yang, S. H. et al. Hum. Mol. Genet. 24, 2826–2840 (2015).
Liao, H. K. et al. Cell 171, 1495–1507 e15 (2017).
Dobin, A. et al. Bioinformatics 29, 15–21 (2013).
Acknowledgements
We thank A. Ocampo for discussions throughout the study and M. Schwarz for administrative support. We are grateful to G. Sancar and Ronald Evans Lab for providing the running wheel equipment. E.B. was partially funded by the Catharina Foundation. P.R. was partially supported by the Muscular Dystrophy Association. G.E. was partially funded by NIH-NCI CCSG: P30 014195 and by the Helmsley Trust. Work in the laboratory of J.C.I.B. was supported by The Progeria Research Foundation, Universidad Católica San Antonio de Murcia (UCAM), Fundacion Dr. Pedro Guillen, the G. Harold and Leila Y. Mathers Charitable Foundation, The Glenn Foundation and The Moxie Foundation.
Author information
Authors and Affiliations
Contributions
E.B., H.-K.L. and J.C.I.B. designed all the experiments. E.B., H.-K.L., P.R. and J.C.I.B. prepared the figures and wrote the manuscript. E.B., H.-K.L. and Y.F. performed and/or analyzed in vitro experiments. E.B., H.-K.L., Y.F., M.Y., R.H.-B. and P.R. performed and/or analyzed ex vivo and in vivo experiments. G.E. performed the bioinformatics analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
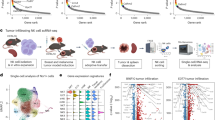
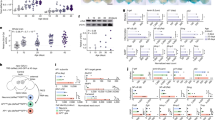
Extended Data Fig. 1 Immunoblots and RNA sequencing of Lmna gRNA-treated fibroblasts of adult HGPS mice.
a, Immunoblots of individual (g1 or g2) or multiplex (g1&2) gRNA-treated fibroblasts derived from adult HGPS (Pro); Cas9 mice. Fibroblasts derived from heterozygous (Pro/+) and homozygous (Pro/Pro) HGPS mice of a heterozygous Cas9 (Cas9/+) background were treated with lentivirally delivered gRNAs. No Cas9 (Pro/+;+/+) sample or mock-treatments were used as negative controls. Middle panels show longer exposures.
Extended Data Fig. 2 Fluorescent images of Lmna gRNA-injected mice.
a, Whole-body fluorescent images of a P4 (4 days dpp, days postpartum) mouse injected at P0 with a 1:1 mix of gRNA1 and gRNA2 viral preparations (each 1.29 × 1011 genomic copies per viral preparation). b, Same organs as in Fig. 1c, under individually adjusted exposure levels relative to negative controls. The organs were harvested from a 6 dpp mouse, 5 days post-injection (dpi) at P1 (P1-5DPI). The inset corresponds to the square area. Ctrl: no injection. c, Expression of the mCherry reporter in the adult (5.5 months post-injection of a 1:1 mix of gRNA1 and 2 viral preparations;1.3 × 1011 genomic copies per viral preparation). The numbers at the upper right corner denote the exposure times. stom, panc, int, sp, kid, br, liv, mscl: stomach, pancreas, intestine, spleen, kidney, brain, liver, muscle, respectively.
Extended Data Fig. 3 Genomic DNA and protein analyses of Lmna gRNA-treated mice.
a, Relative intensities of lamin A, progerin and lamin C bands on the immunoblot shown in Fig. 1e. The intensity of each band was normalized by
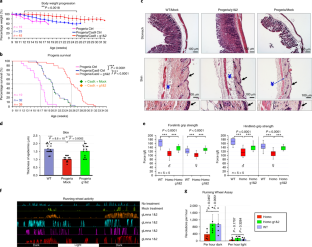
Extended Data Fig. 4 Physiological analyses of Lmna gRNA-treated mice.
a, Histological analysis of the aortic arch of untreated and gRNA-treated (g1&2) HGPS-Cas9 mice, 19 weeks old, injected at P0 with a 1:1 mix of gRNA1 and gRNA2 viral preparations. The black bracket shows the medial layer of the aortic arch. Scale bar, 100
Extended Data Fig. 5 Observed phenotypes of HGPS mice with extended lifespan.
a, Prevalence of megacolon/megacecum among homozygous HGPS mice treated with gLmna-1 and 2 (g1&2) versus negative controls. Asymp: asymptomatic (no obvious gastrointestinal phenotype); G.I: gastrointestinal phenotype characterized by megacolon/megacecum and inability to defecate. b, Chenges in body weight of a representative g1&2-treated homozygous mouse that died suddenly after 28 weeks. c, Necropsy of a g1&2-treated homozygous mouse exhibiting megacolon/megacecum (left and bottom) versus negative control (right).
Supplementary information
Supplementary Information
Supplementary Table 1
Supplementary Video 1
g1&2-treated homozygous mouse (active chinchilla colored) versus its heterozygous sibling (black) and a mock-treated homozygous mouse (inactive chinchilla colored). The treated and heterozygous mice were 19.4 weeks old, 3 weeks older than the mock-treated control. Same animals as in Fig. 1f.
Supplementary Video 2
Same animals as in Video 1, except the black mouse is the mock-treated WT sibling (same as in Fig. 1f) of the negative control.
Source data
Source Data Fig. 1
Source Data (Full Blots)
Rights and permissions
About this article
Cite this article
Beyret, E., Liao, HK., Yamamoto, M. et al. Single-dose CRISPR–Cas9 therapy extends lifespan of mice with Hutchinson–Gilford progeria syndrome. Nat Med 25, 419–422 (2019). https://doi.org/10.1038/s41591-019-0343-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0343-4